Lethbridge High School iGEM
iGEM – Lethbridge
Project Summary
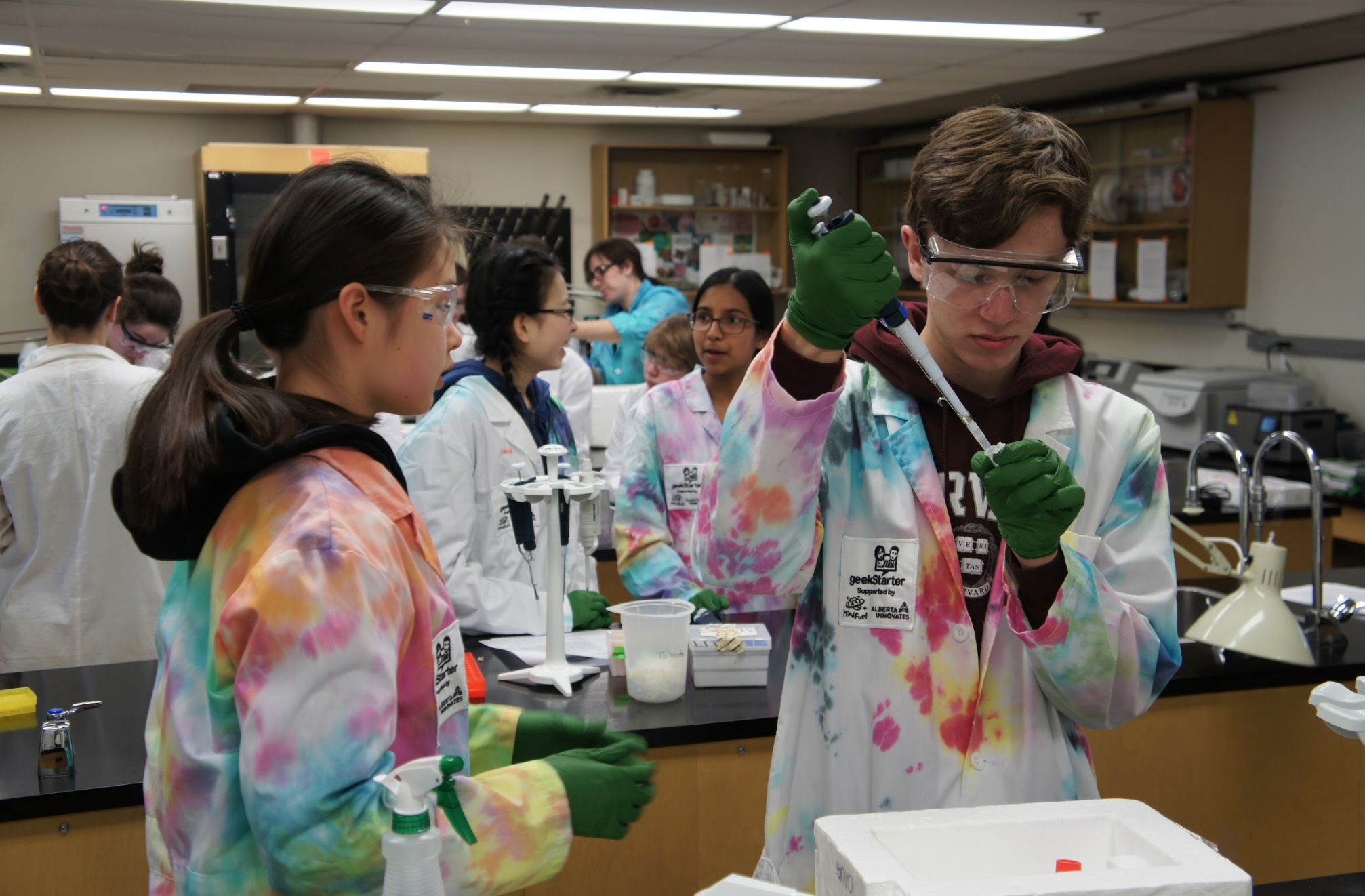
The team has recently completed recruitment for the 2019 iGEM season.the Lethbridge high school team will be working with the CRISPR-Cas13a system to develop a rapid detection system for common hospital and environmental pathogens. They had a tough time deciding on a project as there were so many great options! Currently, the team is working on writing their BioTreks abstract and learning as much about their system as possible. They’re very excited for the upcoming season and can’t wait to get into the brand new lab space!
Project Summary
Last Update: July 2019.
In the coming years, antibiotic-resistant bacteria are expected to become one of the greatest threats to human health. Although antibiotics are credited with improving the health of millions of people since their inception, the overuse of prescription antibiotics has triggered an evolutionary arms race between researchers and pathogens. Additionally, general antibiotics have been shown to cause substantial harm to human microbiomes due to the imbalance of symbiotic and pathogenic bacteria and can result in serious health issues. As such, a modular and specific substitute is sure to replace the current obsolete antibiotics. We propose to use the CRISPR-Cas13a system as a precursor rapid detection and specific targeting system capable of modular design. This system will ideally target a species-specific RNA transcript in the pathogenic bacteria. Once it interacts with the target, Cas13a becomes activated, initiating non-discriminant cleavage of collateral RNA in the environment. This system has been previously shown for its potential in detection. The system will report if the target sequence from a bacteria is present by a measurable fluorescence change due to collateral cleavage of a signal fluorescent RNA. Furthermore, we plan to partner this system with a thoroughly characterized database of RNA target sequences with specific molecular delivery systems. This will allow us to design multiple systems to target a wide array of bacterial threats. Moreover, by measuring co-culture dynamics through our system we can better translate it to an organism model. This will help us understand the system and predict its effects in real-world events. We hope to enable physicians to improve the accuracy of their diagnosis and assist more people in their recovery from bacterial infection.